CaRNetS

CaRNetS: A computational approach that leverages ATAC-seq and RNA-seq datasets to identify drug targets based on tumor lineage. You can find more about this tool in Forbes et al. 2024.

CaRNetS: A computational approach that leverages ATAC-seq and RNA-seq datasets to identify drug targets based on tumor lineage. You can find more about this tool in Forbes et al. 2024.

DGTAC: Machine learning model trained on 3D chromatin conformation data to predict accurate enhancer to gene connections using ATAC-seq and RNA-seq. You can find more about this tool in Xu et al. 2023.
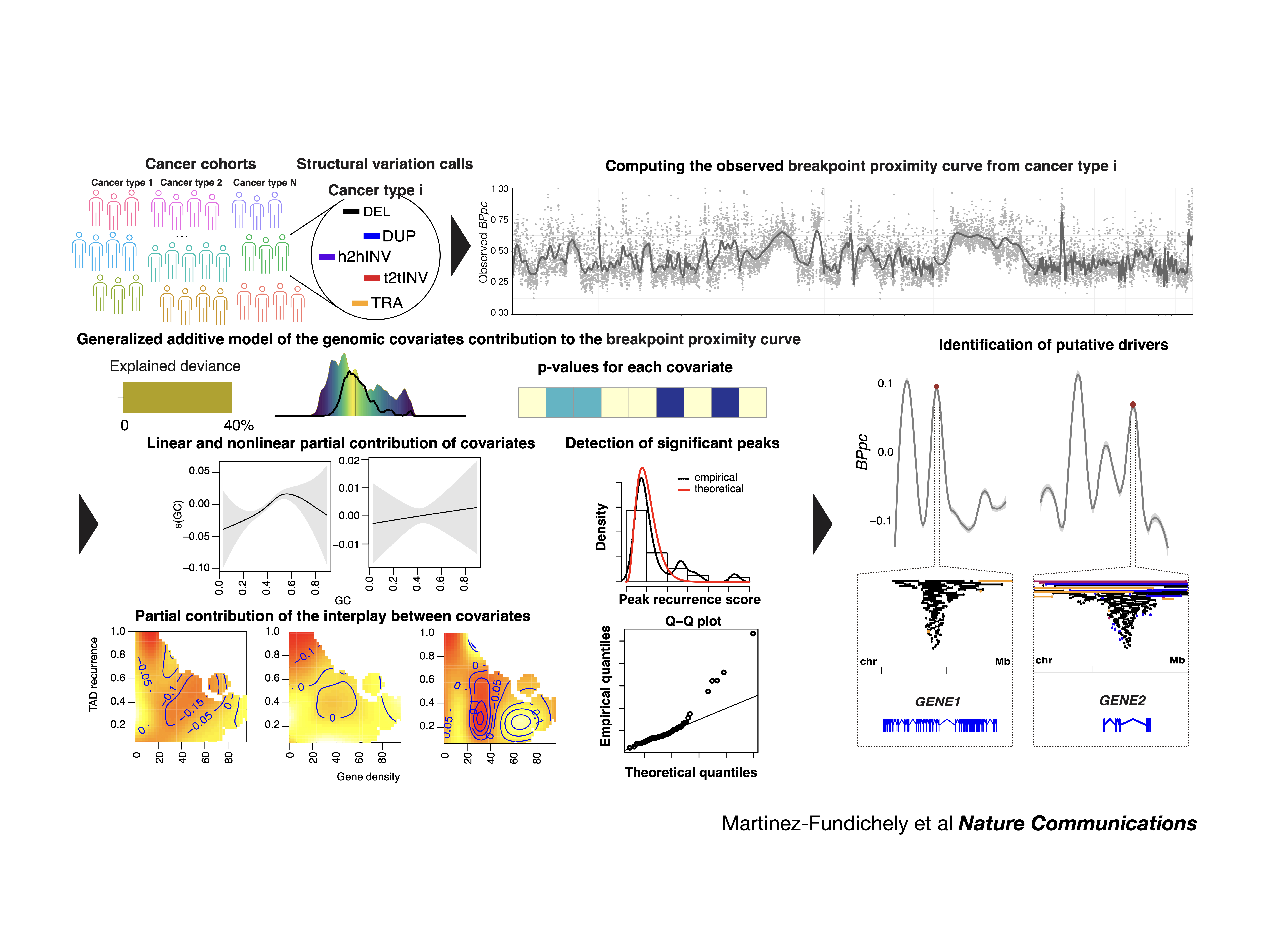
CSVDriver: Identify cancer drivers within significantly rearranged region by modeling tissue-specific breakpoint proximity of structural variations from whole-genomes. You can find more about this tool in Martinez-Fundichely et al. 2022.
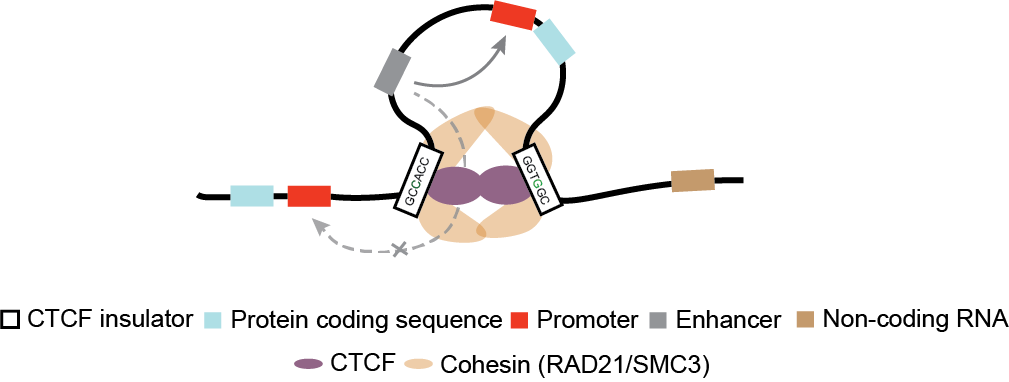
This database provides the list of non-coding cancer drivers in gene promoters, enhancers, ncRNAs and CTCF-cohesin insulators published in over 25 studies. We applied text mining and manual curation to PubMed articles and then developed CNCDatabase, Cornell Non-Coding Cancer driver Database. CNCDatabase provides a helpful resource for researchers to explore the pathological role of non-coding alterations and their associations with gene expression in human cancers. You can also read about it at Liu et al. 2020.
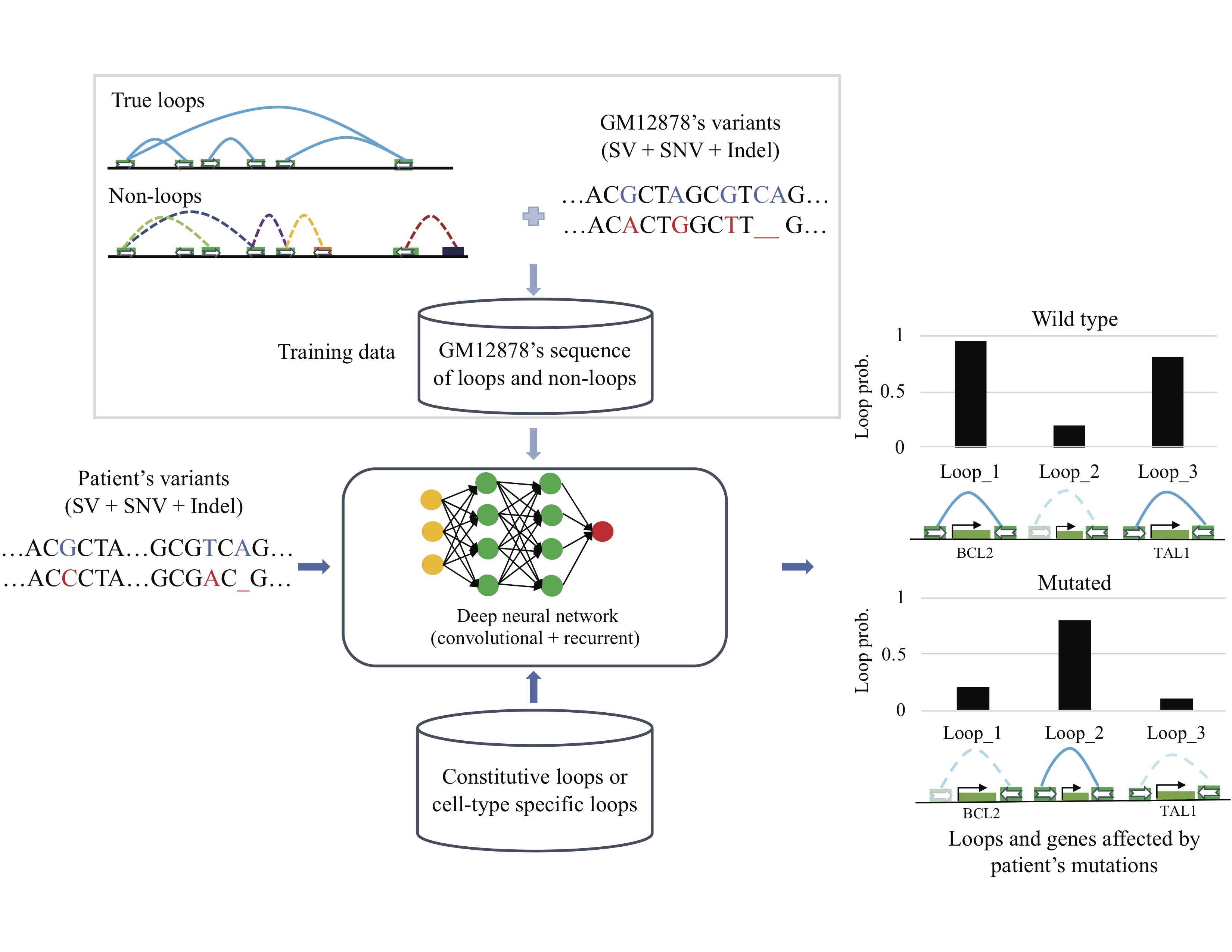
Predicting the impact of non-coding sequence variants on 3D chromatin structure using deep neural network. DeepMILO is available at DeepMILO. You can find more about this tool in Trieu et al. 2020.
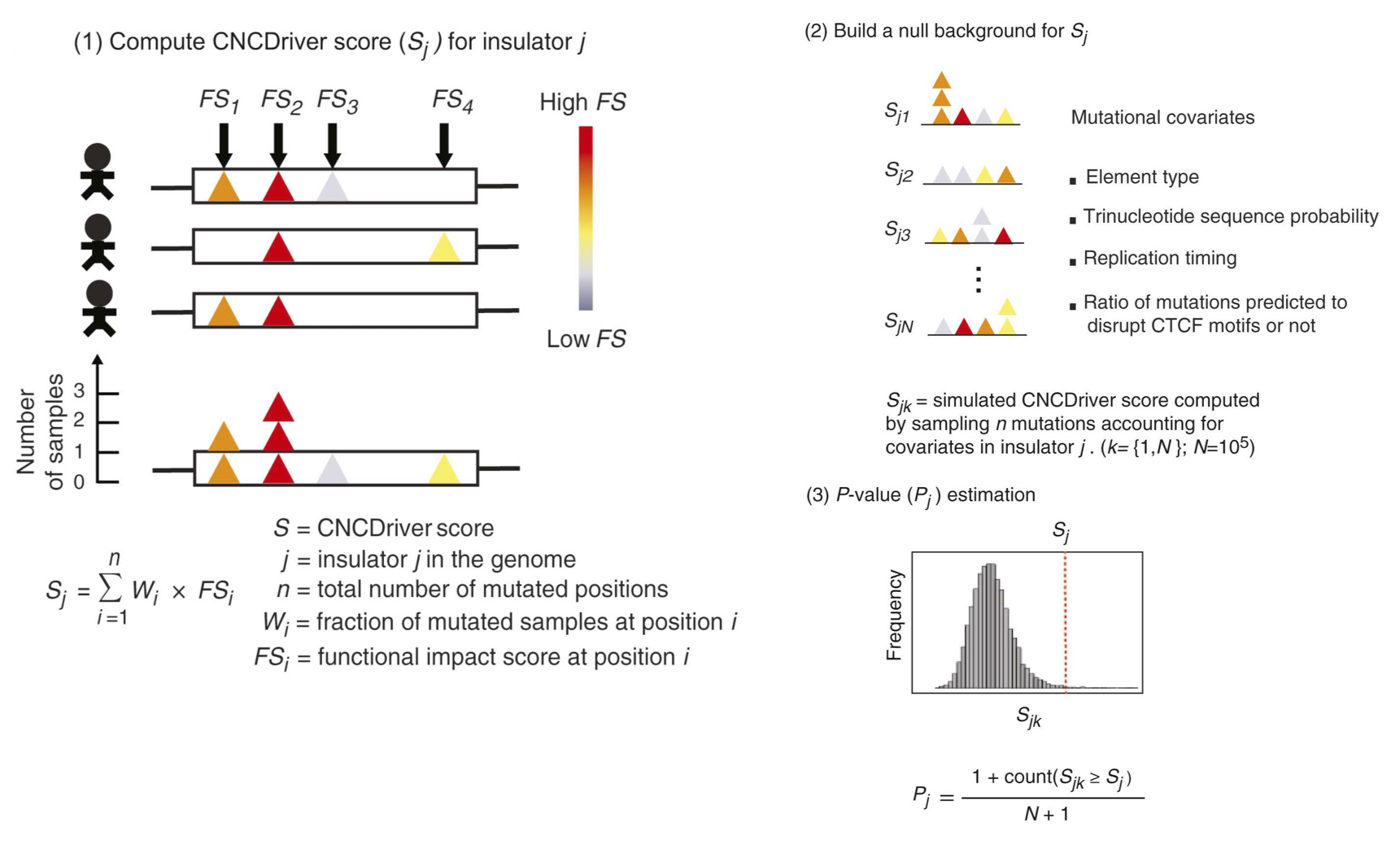
CNCDriver is a method designed to consider mutational recurrence, functional impact and mutational co-variates to identify coding and non-coding cancer driver mutations from whole-genome sequencing data. The source code of CNCDriver is available at CNCDriver, and the details of this method can be found at Liu et al. 2019.
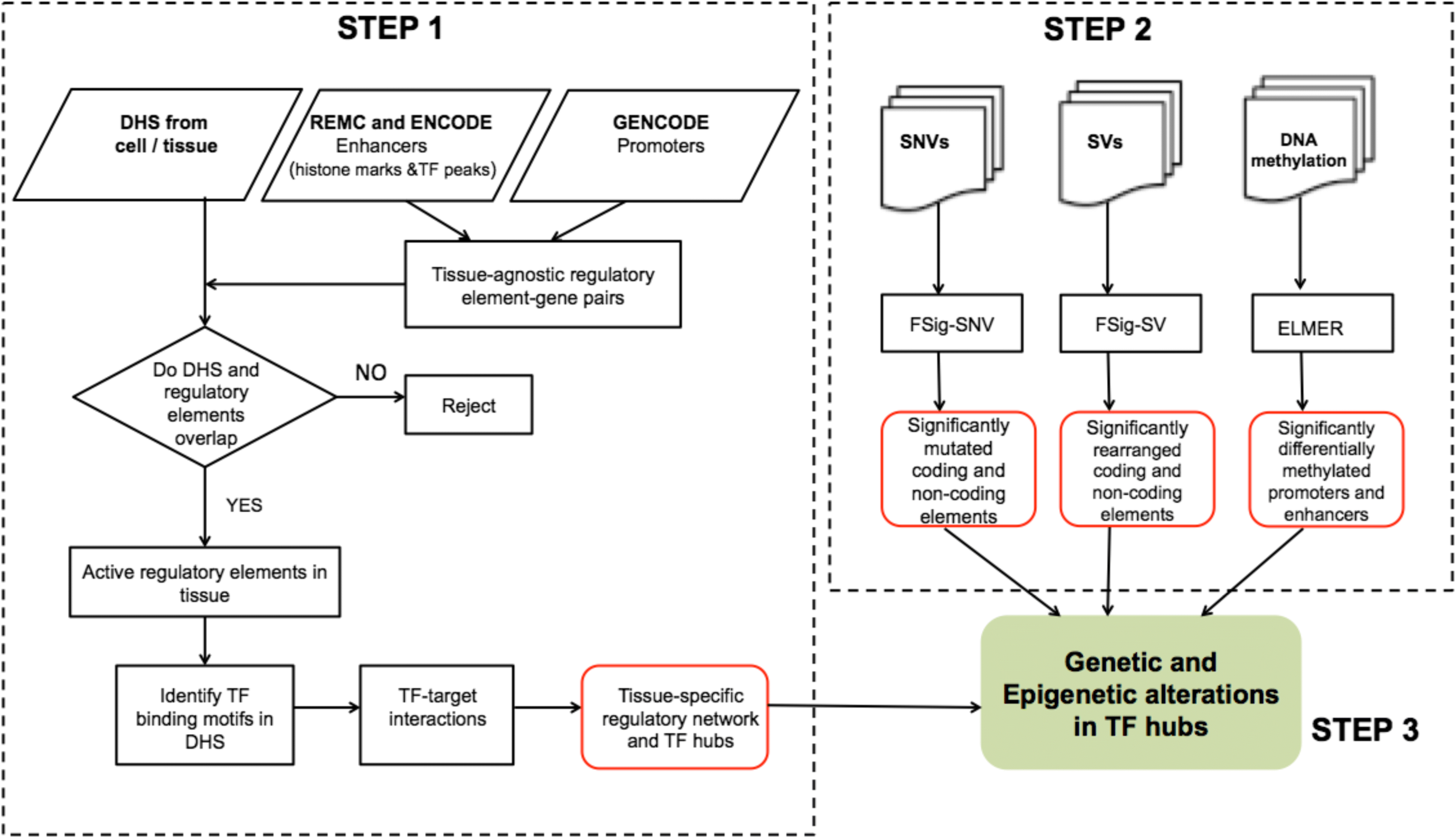
Computational method to identify tumorigenic drivers using the combined effects of coding and non-coding single nucleotide variants, structural variants, and DNA methylation changes in a DNase I hypersensitivity based regulatory network. RegNetDriver is available at RegNetDriver, and more information can be found at Dhingra et al. 2017.
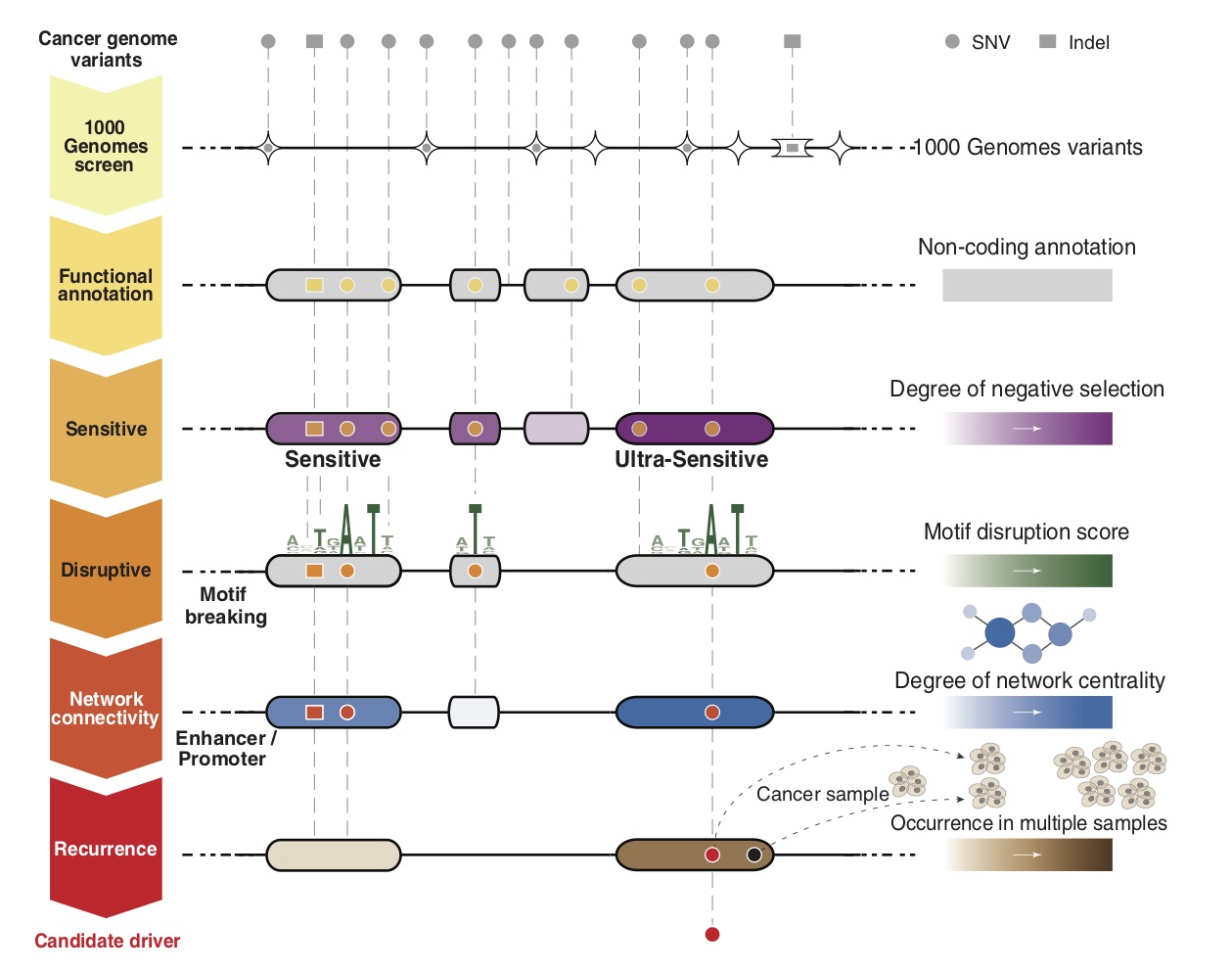
FunSeq2 is a computational framework to annotate and prioritize non-coding drivers from thousands of somatic alterations from cancer whole genomes. FunSeq2 is available at FunSeq2. You can find more about this tool in Fu et al. 2014 and Khurana et al. 2013.